To move toward exascale computing, scientists must focus on saving energy used in computing in every possible way. The energy-conservation requirements go beyond the hardware to each fundamental process of computing, including data management, analysis and visualization. As scientists struggle to manage the massive amount of data generated by experiments and simulations the challenges of data-intensive science and exascale computing begin to merge.
The big-data crush is particularly acute in simulation-dependent science, from high-energy physics and fusion energy to climate, where efforts to analyze and compare models over the next decade will balloon from about 10 petabytes to exabytes.
In today’s high-performance computing (HPC), scientific simulations run on hundreds of thousands of compute cores, generating huge amounts of data that are written on thousands of disk drives or tape. Scientists later process this data, creating new insights.
As simulations and experiments begin to generate extreme amounts of data, new challenges are emerging, especially ones involving energy constraints. To compensate, data management, analysis and visualization processes must be much more integrated with simulations and with each other. As David Rogers, a visualization and data analysis researcher at Sandia National Laboratories, says, “We’re trying to couple more analysis and visualization with simulation so that we can find the most relevant information and make sure that it gets saved.”
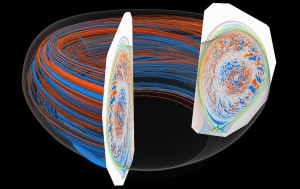

A 3-D visualization of a plasma’s turbulent flow in a tokamak ring helps fusion-energy researchers validate the overall flow structure. The cutaway view allows scientists to study in detail two-dimensional fluctuations and the magnetic separatrix, or the way the device limits the plasma to control the fusion reaction. To fully render and animate data from a simulated turbulent flow and capture its fine details, researchers must develop specialized algorithms that take full advantage of advanced graphics hardware. The complete dataset in a tokamak simulation is large, consisting of several thousands time steps. Interactive visualization would require next-generation supercomputers. Image courtesy of Kwan-Liu Ma and Yubo Zhang, Institute for Ultra-Scale Visualization and University of California-Davis; simulation by C.S. Chang, New York University.
To do that, researchers must build continuous processes into HPC. “We must think of the entire flow, end-to-end knowledge, while designing the system itself, maybe even at the expense of computational capability,” says Alok Choudhary, John G. Searle professor of electrical engineering and computer science at Northwestern University. “We must ask how long it takes to get an answer instead of just how long it takes to do the computation (and) think about extracting data on the fly and analyzing it” as the process runs. This is called the in situ approach.
As HPC evolves toward exascale and data-intensive fields generate exabytes of data , data management, especially moving data, creates an increasingly complicated problem. A critical issue is that the speed of disk storage has not kept pace with increasing processor speeds, while network limitations constrain the speed of input/output (I/O), or the communication between computational components or between the computer and, say, a user or outside system. Also, the power cost of moving data on the computer and especially on and off the computer is very expensive. The problem, Choudhary says, extends from “just thinking about the meaning of doing I/O down to the nuts and bolts of accomplishing that.”
Some existing tools, such as the Adaptable I/O System (ADIOS), show promise for speeding up I/O. Scott Klasky, leader of the scientific data group at Oak Ridge National Laboratory and head of the ADIOS team, says the tool already works with a range of applications, from combustion to fusion to astrophysics. Nonetheless, advances in I/O also must work with many analysis and visualization tools, Klasky says.
Software tools such as ParaView and VisIt – two key Department of Energy visualization platforms – can use the ADIOS framework to run either in situ or on stored data. This enables analysis and visualization without extensive theoretical knowledge. Such tools help enormously with data-intensive science. However, these tools are not expected to scale much beyond today’s resources and will have to be reworked on emerging hardware.
Other approaches also could accelerate I/O. Instead of writing related data to one file, for example, some systems, such as ADIOS, can write faster to multiple subfiles. “This way,” says Arie Shoshani, senior staff scientist at Lawrence Berkeley National Laboratory, “you can write better in parallel and sometimes you get an order of magnitude improvement in I/O.”
‘How can we do exploration in a way that won’t hold the machine hostage?’
But because of stringent energy constraints, researchers will need to reinvent the entire file concept to build storage systems feasible for exascale computing. They must consider alternative approaches to store data, such as solid-state non-volatile random access memory (NVRAM) placed near processors, on the compute nodes. Solid-state memory uses less power than disks and placing memory near processors will reduce the need for time- and energy-consuming data movement. This approach allows scientist to use the deep memory hierarchy – various layers that can be accessed, such as caches – in a consistent programming model for both analytics and simulation and also could enhance I/O. As Choudhary says, “That provides more flexibility to optimize I/O when programming a simulation to access the device.”
New hardware architecture, though, will not solve every issue on the way to exascale computing. “The biggest change will be how scientists think about extracting knowledge from data,” Choudhary says.
To reduce the data being analyzed, researchers concentrate in situ processing on specific features in a simulation. As Shoshani explains: “You might want to look at a bunch of particles moving in accelerators and you can’t see the details in the forest, so maybe you could concentrate on particles with a certain energy or momentum.” Then, the software would need to quickly reduce the data for those particles. “Indexing the data comes in especially useful if doing this interactively.”
Maybe tomorrow’s simulations will analyze chunks of data as they are generated. Imagine a simulation producing data across time steps. Perhaps, as a very simple idea, a statistical procedure could reduce the data from several steps to one.
More complex problems arise when mining data generated in exascale machines. As Hank Childs, a computer systems engineer at Lawrence Berkeley National Laboratory, asks, “How can we do exploration in a way that won’t hold the machine hostage?”
How the data are stored also will affect later data-mining options. Wes Bethel, senior computer scientist at Lawrence Berkeley National Laboratory, explains this with a climate-model scenario. Imagine a grid that represents some area, and each point in the grid includes, say, a dozen variables, such as temperature, pressure and so on. A simulation typically generates the data for a grid point and then stores the results for each variable. Then it moves to the next grid point and stores those data. If a researcher wants to look at temperature over a century, the program first must seek the temperature variable at grid-point 1 at time-step 1 and then do that for each grid point and time step. “It takes forever to read these data,” Bethel says. “If all of the temperature variables were together, then the downstream analysis process would be more efficient.”
Test runs on computers with lower performance will help researchers know what to look for even in the unknown. As Rogers points out, “An exascale simulation is not something that you think up one day and do the next.” Instead, researchers start with smaller-scale projects and work up to exascale, studying each simulation in detail. This helps them identify areas of the exascale simulation that are likely to be interesting so they can focus their analysis on those.
Over the history of HPC, advances focused on computers churning out the desired computations. “We need that, and it’s critical,” says Sean Ahern, computer scientist at Oak Ridge National Laboratory, “but solving mathematical equations that describe physical reality is only half the job.” The other half, Ahern says, is visualization – “taking information and turning it into images that can be understood.”
Increasing the complexity of computing does not necessarily mean more complex visualization. “I have seen movies published at visualization conferences, and colleagues look at them and say, ‘It looks great, but did it tell you anything about the science?’” Ahern says. The “best visualization could be as simple as a number, say, 12.7, or a really complex volume rendering. It’s scientific understanding that we really care about.”
To get the best visualization as computation moves to the exascale, researchers must look ahead. “The most successful projects think about how to understand data from the get-go,” Ahern says, though it’s “moderately rare to do that.” In exascale machines, researchers will need to know how data will be displayed and how the displays will be generated on the fly.
Exascale simulations also will produce too much data, a problem that already exists to a lesser degree. “Even today, the scientists we work with can store only a small fraction of the data generated from simulations,” says Kwan-Liu Ma, professor of computer science at the University of California, Davis. “Maybe a simulation runs 100,000 time steps, and the scientists store only, say, every 300th.” Moving toward exascale, scientists will store even less of the data generated and next-generation experimental facilities will also have to grapple with some sort of data triage as multiple petabytes of data poor from detectors every second. With a smaller percentage of the data stored, an important challenge is how to choose data that will be stored so that exploratory analysis still permits discovery of the unexpected.
So visualization scientists must collaborate with experiment and simulation teams more than ever. Ma describes a group effort with Sandia scientists on a direct numerical simulation of turbulence in combustion. “We tried to tightly couple the visualization code with their simulation code to capture a snapshot of the data at the highest possible resolution before reduction,” Ma says, “but that’s not enough.” Beyond coupling the processes, the visualization code must be just as scalable as the simulation code or the visualization will slow down the entire process.
In some cases, researchers will continue to rely on reduced data that are visualized later. As Ma says, though, the data would have to be prepared “in the right way during the simulation because it would be too expensive to do that after the simulation.”
To plan the combination of data management, analysis and visualization with exascale computers and next-generation experimental facilities, researchers will need to understand the hardware and architecture of those machines, and that is evolving. Consequently, speculation still surrounds the best approaches to working with the data and getting the most out of data-intensive science.