Bert Debusschere digs around in some messy stuff. He looks for ways to understand chemical and biochemical reactions that are highly variable, hard to distinguish from background activity and often sensitive to small changes in conditions. The data rarely are neat and clean; instead, they’re consistently inconsistent.

Bert Debusschere
Nonetheless, they’re important reactions – parts of networks governing gene regulation, the human immune system and catalysis in fuel cells and batteries. If we can decipher these stochastic reaction networks, there could be big rewards, such as improved energy efficiency, new disease treatments and better environmental remediation methods.
Debusschere, a staff member at Sandia National Laboratory’s Transportation Energy Center in Livermore, Calif., develops computational and mathematical methods to study these hard-to-characterize systems. For his work, Debusschere was one of 68 researchers to receive the 2007 Presidential Early Career Award for Scientists and Engineers, presented at a White House ceremony in December 2008.
The award also recognizes Debusschere’s work with the Sandia Diversity Council and Foreign National Networking Group – but omits the nonprofit canine rescue operation that takes up most of his off-work hours.
Debusschere’s group already is working with a biomedical researcher to apply its methods to immune system signaling pathways. But, he says: “The work we’re doing is really on a very fundamental methodology level. We’re probably a couple years away from having this in every biochemist’s toolbox, but it’s making its way over there.”
Debusschere received his bachelor’s degree in mechanical engineering in his native Belgium before earning master’s and doctoral degrees in the same subject at the University of Wisconsin. He specialized in computational fluid dynamics with direct numerical simulations of scalar transport in reacting flows.
Such stochastic reaction networks are everywhere, Debusschere says, making the tools he and his fellow researchers develop broadly applicable.
When he first arrived at Sandia in 2001, Debusschere researched ways to assess the uncertainty in models of the fluid flows behind combustion and similar processes. On the large scale, so many reactions occur in these systems that the models are mostly deterministic, with a clearly defined path, and scientists have good tools for analyzing them.
Debusschere soon became intrigued by biological system models, which can confound tools used to analyze uncertainty in deterministic models.
There are two main problems: First, the scale at which these reactions take place is so small that even those involving individual molecules generate great variability or “noise.” “The fact they’re so tiny makes them hard to resolve discriminately. You’re counting molecules rather than concentration levels,” Debusschere says.
Second, experiments generally capture but a sparse subset of molecules reacting in the system.
The noise in the reactions creates intrinsic uncertainty, which contributes to model uncertainty: Given the variability in such systems, how can researchers even be sure they have the right model for what they’re studying?
With a standard deterministic model, “every time you run it with the same parameter you get the same answer,” Debusschere says. “With intrinsic variability, every time you run the model you can get a different answer,” even with identical parameters – and each answer is valid.
Finally, there’s also parametric uncertainty – will a model make sense, given a certain set of parameters that influence the process? Different reaction networks are more or less sensitive to parameter changes, and it’s often difficult to sort them out.
Because of intrinsic variability, it’s difficult to compare predictions with experiments or to infer a model that could explain experiments, Debusschere says. Researchers try to understand what parameters are most responsible for an outcome or change the parameters to produce a desired result, but they often can’t do either with much confidence.
“People have come a long way in figuring out these models, but what’s missing is a comprehensive computer framework to investigate the uncertainties,” he says.
That’s Debusschere’s mission. With Khachik Sargsyan and Habib Najm at Sandia, Olivier Le Maître of France’s Laboratoire d’Informatique pour la Mécanique et les Sciences de l’Ingénieur and others, he’s developing tools to analyze intrinsic noise and parametric uncertainty.
It starts with probability. In stochastic reaction networks, each time point presents “a certain probability of reactions taking place rather than a certain rate of reactions taking place,” Debusschere says.
“You have to properly define what your observable is,” he adds – what molecule or reaction you want to understand. But “you can’t say ‘I’m interested in the number of molecules of X.’ You will get a different answer each time, so you look at a distribution of the number of molecules.” By running models multiple times, the researchers can get a probability distribution of reaction products.
Debusschere’s main tools for understanding uncertainties are polynomial chaos expansions (PCEs). Polynomial chaos methods, developed in the 1930s by American mathematician Norbert Wiener, are a way to reduce the complexity of random variables, making them easier to work with in computations. They provide compact representations of how intrinsic noise and parametric uncertainty affect a system, making the corresponding processes easier to analyze.
“PCEs present a way to describe stochasticity with a set of deterministic coefficients,” Debusschere says. “Those deterministic descriptions are a lot easier to work with.”
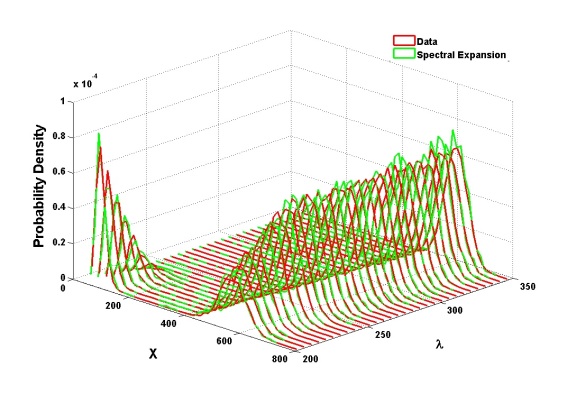
In most cases, Debusschere and his fellow researchers have used PCEs to represent different types of variability and uncertainties. Their standard test case has been the Schlögl model, a prototype reaction network in which two chemical concentrations are stable while concentrations of other chemicals, reaction rates and other parameters change. Their PCE-based methods demonstrate good agreement with original data that show the system moving from two-activity modes to a single mode as values of a selected parameter increase.
In one case, Debusschere and his fellow researchers used PCEs to artificially perturb parameters to learn about the system’s sensitivity to such changes. The researchers gave selected parameters a quantified uncertainty large enough to generate a response that would stick out over intrinsic noise. On a test using a simplified virus infection model, the researchers found the probability of failed infections, the viral reservoir (cells where the infection is latent for a long time without fully developing) and virus production rate all are very sensitive to the generation of two particular viral nucleic acids.
Such stochastic reaction networks are everywhere, Debusschere says, making the tools he and his fellow researchers develop broadly applicable. They can be used to study the interactions and feedback loops governing cell growth, gene expression and other biological activities. They can study catalytic reactions important for batteries and fuel cells. They can help understand the conditions affecting how bacteria break down environmental contaminants or plant material so scientists can tweak the bacteria and conditions to boost efficiency.
In perhaps the most promising application to date, Debusschere’s group is teaming with researchers at the University of Texas Medical Branch (UTMB) in Galveston. Led by Dr. Allan Brasier, a professor in the Department of Internal Medicine and Institute of Translational Sciences, the UTMB group is investigating the causes of inflammation, an underlying mechanism in diseases such as atherosclerotic heart disease and asthma. They hope to sift out the cell signaling paths inflammation activates. If researchers can decipher these signaling pathways, they may be able to modify them to develop new treatments.
The team has focused on cell signal pathways that govern nuclear factor-κB (NF-κB). NF-κB, a protein complex that regulates the innate immune response, enters the cell nucleus and binds to regulatory DNA sequences. This triggers expression of a network of inflammatory genes, says Brasier, who also is director of UTMB’s Sealy Center for Molecular Medicine.
The pathways controlling NF-κB are complex, Brasier says, and can be induced by a number of different substances, including viruses and regulatory proteins and peptides. “All of these pathways activate different subsets of the NF-κB proteins with different kinetics, and are mediated by different intracellular signaling cascades,” he says. How much each cell activates NF-κB and responds to it can vary significantly. This variation may have important consequences for the evolution of the inflammatory response.
Brasier says his group already uses biochemical approaches to determine the pathway structures. “Where Bert will have a major impact is to identify the key steps regulating those pathways,” he adds.
With a pilot grant from a joint UTMB-Sandia institute, “We’re in the process of collecting data and evaluating published models,” Brasier says. “Right now, Bert is focusing on identifying which parameters in the models are most important that need to have additional measurements made. This is a repeating process of modeling, experimental observations, refining the models, et cetera.”
This type of dynamic understanding of the pathway behavior isn’t possible without Debusschere’s computational insight, Brasier says.
The methods Debusschere and his fellow researchers have developed also could soon see wider use through an uncertainty quantification toolkit. The methods could help researchers understand how much confidence they can have in results from their computational models.
“We’re working on putting together a toolkit that gathers all these things and makes them available to the general user in the (computational science) community,” Debusschere says. The group hopes to release it later in 2009 as open-source software.
“The toolkit is something we have worked on with a lot of people in this area,” he says. “It’s not the product of one specific project but it has contributions from every project that used PCEs.”